Although DNA DNA A deoxyribonucleotide polymer that is the primary genetic material of all cells. Eukaryotic and prokaryotic organisms normally contain DNA in a double-stranded state, yet several important biological processes transiently involve single-stranded regions. DNA, which consists of a polysugar-phosphate backbone possessing projections of purines (adenine and guanine) and pyrimidines (thymine and cytosine), forms a double helix that is held together by hydrogen bonds between these purines and pyrimidines (adenine to thymine and guanine to cytosine). DNA Types and Structure fidelity Fidelity Conflict of Interest is highly protected, DNA DNA A deoxyribonucleotide polymer that is the primary genetic material of all cells. Eukaryotic and prokaryotic organisms normally contain DNA in a double-stranded state, yet several important biological processes transiently involve single-stranded regions. DNA, which consists of a polysugar-phosphate backbone possessing projections of purines (adenine and guanine) and pyrimidines (thymine and cytosine), forms a double helix that is held together by hydrogen bonds between these purines and pyrimidines (adenine to thymine and guanine to cytosine). DNA Types and Structure can still be damaged by a number of environmental factors, reactive oxygen species Reactive oxygen species Molecules or ions formed by the incomplete one-electron reduction of oxygen. These reactive oxygen intermediates include singlet oxygen; superoxides; peroxides; hydroxyl radical; and hypochlorous acid. They contribute to the microbicidal activity of phagocytes, regulation of signal transduction and gene expression, and the oxidative damage to nucleic acids; proteins; and lipids. Metabolic Dysfunction-associated Steatotic Liver Disease (MASLD), and errors in DNA replication DNA replication The entire DNA of a cell is replicated during the S (synthesis) phase of the cell cycle. The principle of replication is based on complementary nucleotide base pairing: adenine forms hydrogen bonds with thymine (or uracil in RNA) and guanine forms hydrogen bonds with cytosine. DNA Replication. DNA DNA A deoxyribonucleotide polymer that is the primary genetic material of all cells. Eukaryotic and prokaryotic organisms normally contain DNA in a double-stranded state, yet several important biological processes transiently involve single-stranded regions. DNA, which consists of a polysugar-phosphate backbone possessing projections of purines (adenine and guanine) and pyrimidines (thymine and cytosine), forms a double helix that is held together by hydrogen bonds between these purines and pyrimidines (adenine to thymine and guanine to cytosine). DNA Types and Structure repair is a continuous process in which the cell corrects the damage. The cell has multiple mechanisms it can use to repair DNA DNA A deoxyribonucleotide polymer that is the primary genetic material of all cells. Eukaryotic and prokaryotic organisms normally contain DNA in a double-stranded state, yet several important biological processes transiently involve single-stranded regions. DNA, which consists of a polysugar-phosphate backbone possessing projections of purines (adenine and guanine) and pyrimidines (thymine and cytosine), forms a double helix that is held together by hydrogen bonds between these purines and pyrimidines (adenine to thymine and guanine to cytosine). DNA Types and Structure. During replication, the cell has proofreading machinery within the DNA DNA A deoxyribonucleotide polymer that is the primary genetic material of all cells. Eukaryotic and prokaryotic organisms normally contain DNA in a double-stranded state, yet several important biological processes transiently involve single-stranded regions. DNA, which consists of a polysugar-phosphate backbone possessing projections of purines (adenine and guanine) and pyrimidines (thymine and cytosine), forms a double helix that is held together by hydrogen bonds between these purines and pyrimidines (adenine to thymine and guanine to cytosine). DNA Types and Structure polymerase itself. For single-stranded DNA DNA A deoxyribonucleotide polymer that is the primary genetic material of all cells. Eukaryotic and prokaryotic organisms normally contain DNA in a double-stranded state, yet several important biological processes transiently involve single-stranded regions. DNA, which consists of a polysugar-phosphate backbone possessing projections of purines (adenine and guanine) and pyrimidines (thymine and cytosine), forms a double helix that is held together by hydrogen bonds between these purines and pyrimidines (adenine to thymine and guanine to cytosine). DNA Types and Structure damage, the cell can use excision repair techniques and photorepair. For double-stranded DNA DNA A deoxyribonucleotide polymer that is the primary genetic material of all cells. Eukaryotic and prokaryotic organisms normally contain DNA in a double-stranded state, yet several important biological processes transiently involve single-stranded regions. DNA, which consists of a polysugar-phosphate backbone possessing projections of purines (adenine and guanine) and pyrimidines (thymine and cytosine), forms a double helix that is held together by hydrogen bonds between these purines and pyrimidines (adenine to thymine and guanine to cytosine). DNA Types and Structure breaks, the cell can use homologous recombination Recombination Production of new arrangements of DNA by various mechanisms such as assortment and segregation, crossing over; gene conversion; genetic transformation; genetic conjugation; genetic transduction; or mixed infection of viruses. Virology or nonhomologous end joining. When normal DNA DNA A deoxyribonucleotide polymer that is the primary genetic material of all cells. Eukaryotic and prokaryotic organisms normally contain DNA in a double-stranded state, yet several important biological processes transiently involve single-stranded regions. DNA, which consists of a polysugar-phosphate backbone possessing projections of purines (adenine and guanine) and pyrimidines (thymine and cytosine), forms a double helix that is held together by hydrogen bonds between these purines and pyrimidines (adenine to thymine and guanine to cytosine). DNA Types and Structure repair processes fail owing to age, dysfunction, or an overloaded system, unrepaired DNA DNA A deoxyribonucleotide polymer that is the primary genetic material of all cells. Eukaryotic and prokaryotic organisms normally contain DNA in a double-stranded state, yet several important biological processes transiently involve single-stranded regions. DNA, which consists of a polysugar-phosphate backbone possessing projections of purines (adenine and guanine) and pyrimidines (thymine and cytosine), forms a double helix that is held together by hydrogen bonds between these purines and pyrimidines (adenine to thymine and guanine to cytosine). DNA Types and Structure damage can lead to apoptosis Apoptosis A regulated cell death mechanism characterized by distinctive morphologic changes in the nucleus and cytoplasm, including the endonucleolytic cleavage of genomic DNA, at regularly spaced, internucleosomal sites, I.e., DNA fragmentation. It is genetically-programmed and serves as a balance to mitosis in regulating the size of animal tissues and in mediating pathologic processes associated with tumor growth. Ischemic Cell Damage, cellular senescence, or malignant tumors.
Last updated: Dec 15, 2025
DNA DNA A deoxyribonucleotide polymer that is the primary genetic material of all cells. Eukaryotic and prokaryotic organisms normally contain DNA in a double-stranded state, yet several important biological processes transiently involve single-stranded regions. DNA, which consists of a polysugar-phosphate backbone possessing projections of purines (adenine and guanine) and pyrimidines (thymine and cytosine), forms a double helix that is held together by hydrogen bonds between these purines and pyrimidines (adenine to thymine and guanine to cytosine). DNA Types and Structure damage can be caused by:
Proofreading occurs during DNA replication DNA replication The entire DNA of a cell is replicated during the S (synthesis) phase of the cell cycle. The principle of replication is based on complementary nucleotide base pairing: adenine forms hydrogen bonds with thymine (or uracil in RNA) and guanine forms hydrogen bonds with cytosine. DNA Replication. DNA DNA A deoxyribonucleotide polymer that is the primary genetic material of all cells. Eukaryotic and prokaryotic organisms normally contain DNA in a double-stranded state, yet several important biological processes transiently involve single-stranded regions. DNA, which consists of a polysugar-phosphate backbone possessing projections of purines (adenine and guanine) and pyrimidines (thymine and cytosine), forms a double helix that is held together by hydrogen bonds between these purines and pyrimidines (adenine to thymine and guanine to cytosine). DNA Types and Structure polymerase (the enzyme complex that replicates DNA DNA A deoxyribonucleotide polymer that is the primary genetic material of all cells. Eukaryotic and prokaryotic organisms normally contain DNA in a double-stranded state, yet several important biological processes transiently involve single-stranded regions. DNA, which consists of a polysugar-phosphate backbone possessing projections of purines (adenine and guanine) and pyrimidines (thymine and cytosine), forms a double helix that is held together by hydrogen bonds between these purines and pyrimidines (adenine to thymine and guanine to cytosine). DNA Types and Structure):
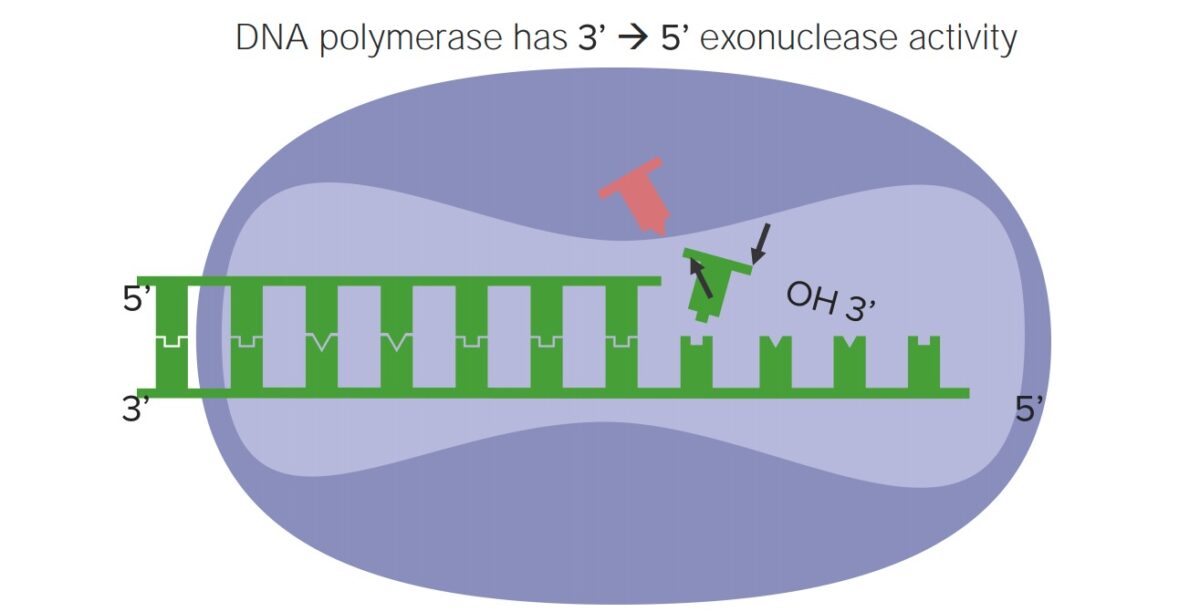
Proofreading activity of DNA polymerase
Image by Lecturio.UV light UV light That portion of the electromagnetic spectrum immediately below the visible range and extending into the x-ray frequencies. The longer wavelengths (near-uv or biotic or vital rays) are necessary for the endogenous synthesis of vitamin D and are also called antirachitic rays; the shorter, ionizing wavelengths (far-uv or abiotic or extravital rays) are viricidal, bactericidal, mutagenic, and carcinogenic and are used as disinfectants. Bullous Pemphigoid and Pemphigus Vulgaris radiation Radiation Emission or propagation of acoustic waves (sound), electromagnetic energy waves (such as light; radio waves; gamma rays; or x-rays), or a stream of subatomic particles (such as electrons; neutrons; protons; or alpha particles). Osteosarcoma causes pyrimidine (T or C) dimers to form via covalent linkages between adjacent bases Bases Usually a hydroxide of lithium, sodium, potassium, rubidium or cesium, but also the carbonates of these metals, ammonia, and the amines. Acid-Base Balance, creating a conformational change (“bulge”) in the DNA DNA A deoxyribonucleotide polymer that is the primary genetic material of all cells. Eukaryotic and prokaryotic organisms normally contain DNA in a double-stranded state, yet several important biological processes transiently involve single-stranded regions. DNA, which consists of a polysugar-phosphate backbone possessing projections of purines (adenine and guanine) and pyrimidines (thymine and cytosine), forms a double helix that is held together by hydrogen bonds between these purines and pyrimidines (adenine to thymine and guanine to cytosine). DNA Types and Structure. These defects can be repaired via a process called photorepair, or photoreactivation.
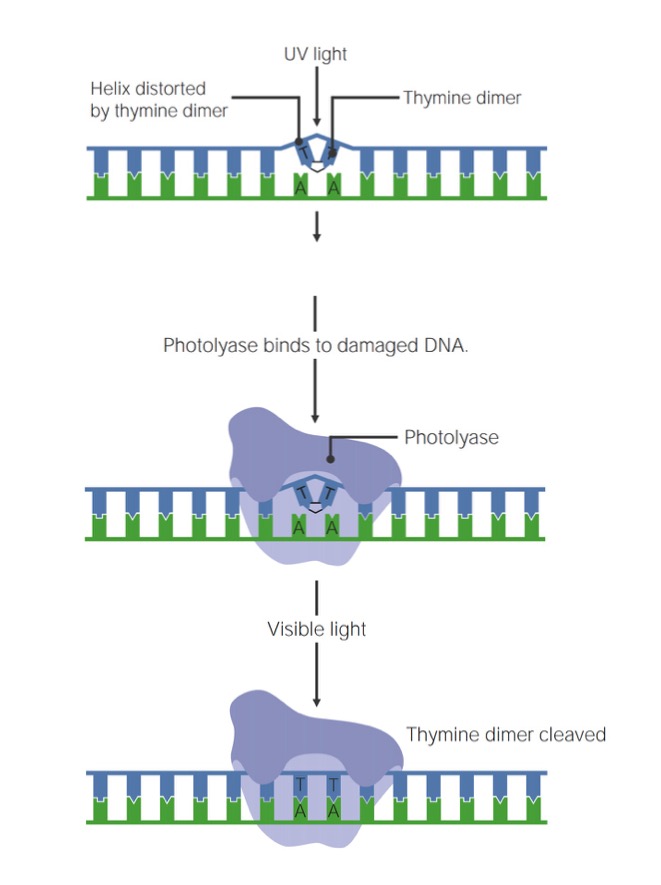
Steps of photorepair
Image by Lecturio.The cell has 3 primary mechanisms to repair damage to a single strand of DNA DNA A deoxyribonucleotide polymer that is the primary genetic material of all cells. Eukaryotic and prokaryotic organisms normally contain DNA in a double-stranded state, yet several important biological processes transiently involve single-stranded regions. DNA, which consists of a polysugar-phosphate backbone possessing projections of purines (adenine and guanine) and pyrimidines (thymine and cytosine), forms a double helix that is held together by hydrogen bonds between these purines and pyrimidines (adenine to thymine and guanine to cytosine). DNA Types and Structure:
All 3 mechanisms follow the same general process:
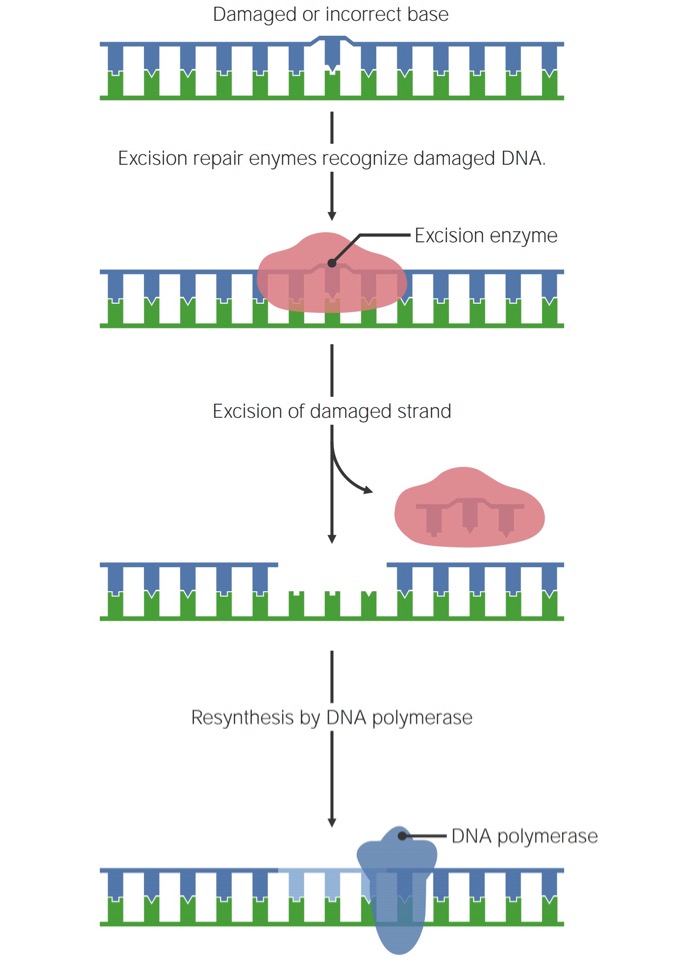
General mechanism of single-stranded DNA repair
Image by Lecturio.In BER, a single damaged base is excised and replaced.
In general, double-stranded DNA DNA A deoxyribonucleotide polymer that is the primary genetic material of all cells. Eukaryotic and prokaryotic organisms normally contain DNA in a double-stranded state, yet several important biological processes transiently involve single-stranded regions. DNA, which consists of a polysugar-phosphate backbone possessing projections of purines (adenine and guanine) and pyrimidines (thymine and cytosine), forms a double helix that is held together by hydrogen bonds between these purines and pyrimidines (adenine to thymine and guanine to cytosine). DNA Types and Structure damage is harder to repair because there is no template strand to work off of. The 2 primary mechanisms to fix double-stranded DNA DNA A deoxyribonucleotide polymer that is the primary genetic material of all cells. Eukaryotic and prokaryotic organisms normally contain DNA in a double-stranded state, yet several important biological processes transiently involve single-stranded regions. DNA, which consists of a polysugar-phosphate backbone possessing projections of purines (adenine and guanine) and pyrimidines (thymine and cytosine), forms a double helix that is held together by hydrogen bonds between these purines and pyrimidines (adenine to thymine and guanine to cytosine). DNA Types and Structure breaks are homologous recombination Recombination Production of new arrangements of DNA by various mechanisms such as assortment and segregation, crossing over; gene conversion; genetic transformation; genetic conjugation; genetic transduction; or mixed infection of viruses. Virology and nonhomologous end joining.
In homologous recombination Recombination Production of new arrangements of DNA by various mechanisms such as assortment and segregation, crossing over; gene conversion; genetic transformation; genetic conjugation; genetic transduction; or mixed infection of viruses. Virology (HR), the nearly identical sister chromatid or homologous chromosome Chromosome In a prokaryotic cell or in the nucleus of a eukaryotic cell, a structure consisting of or containing DNA which carries the genetic information essential to the cell. Basic Terms of Genetics is used as a template:
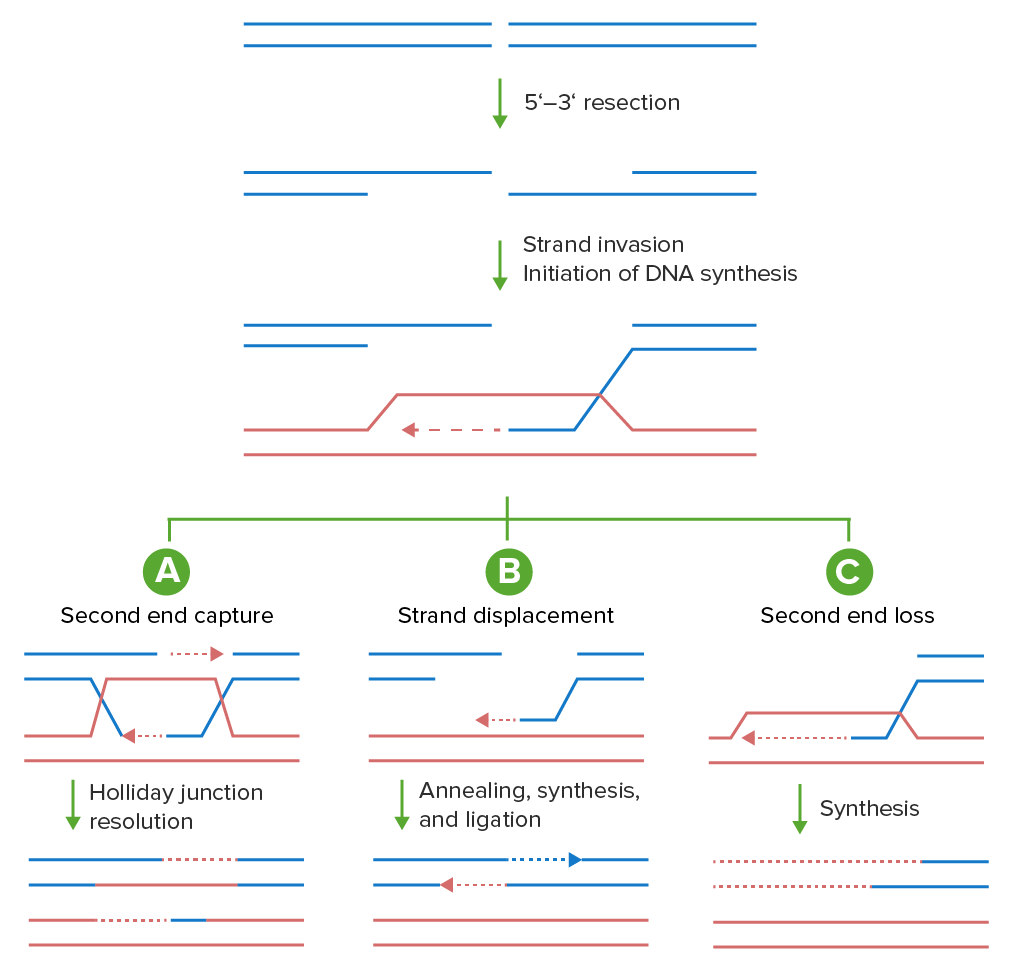
Models of homologous recombination:
Double-stranded breaks can be repaired using the homologous recombination machinery in a variety of ways. The DNA ends are first processed into 3′ single-stranded DNA tails. These tails invade a homologous template (red), priming new DNA synthesis (dashed line). Shown are 3 possible outcomes from this invasion.
A: In canonical double-stranded break repair (DSBR), both the initial invading strand and the captured second end anneal to the homologous template and prime new DNA synthesis, resulting in a double Holliday junction that can be resolved by nucleases into a crossover or a noncrossover product (noncrossover product shown).
B: Alternatively, after the single-stranded DNA tail invades the homologous template, a round of DNA synthesis is prepared from the 3′ end (dashed red line). Synthesis-dependent strand annealing (SDSA) occurs when the invading strand, along with the newly synthesized segment, is unwound by a helicase and annealed with the other resected end.
C: In break-induced replication (BIR), 1 end of the DSB is lost and the remaining end invades the homologous template priming DNA synthesis to the end of the chromosome.