The entire DNA of a cell is replicated during the S (synthesis) phase of the cell cycle. The principle of replication is based on complementary nucleotide base pairing: adenine forms hydrogen bonds with thymine (or uracil in RNA) and guanine forms hydrogen bonds with cytosine. Replication occurs prior to cell division in the S phase of the cell cycle in order to allow 2 sets of chromosomes to be present during the metaphase of mitosis, after which they are equally partitioned into 2 new cells during separation (anaphase).
Last updated: Apr 23, 2025
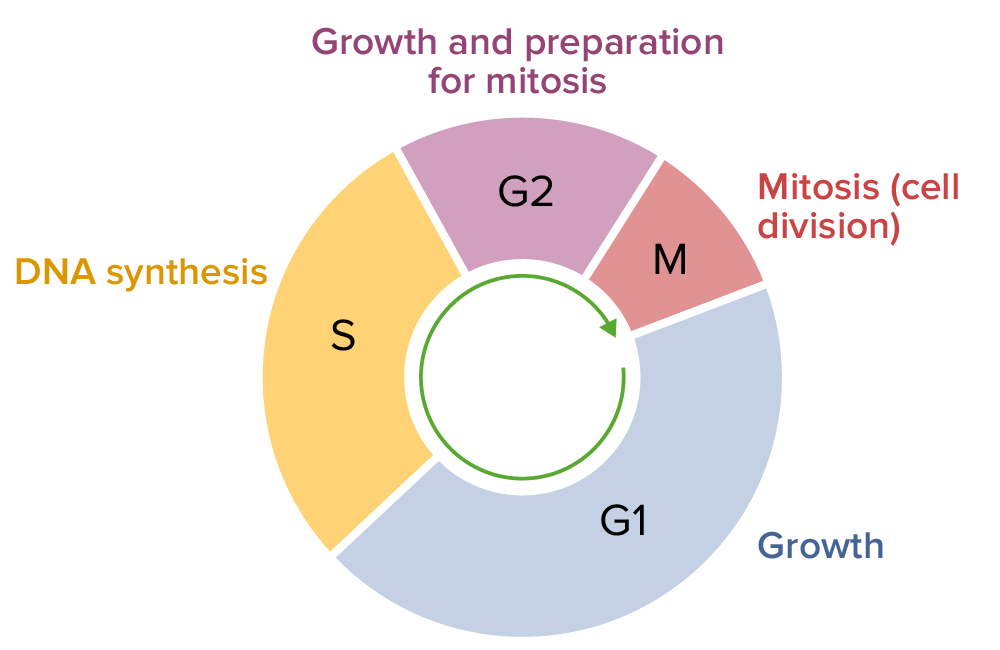
Cell cycle, consisting of G1, S, G2, and M phases
Image by Lecturio.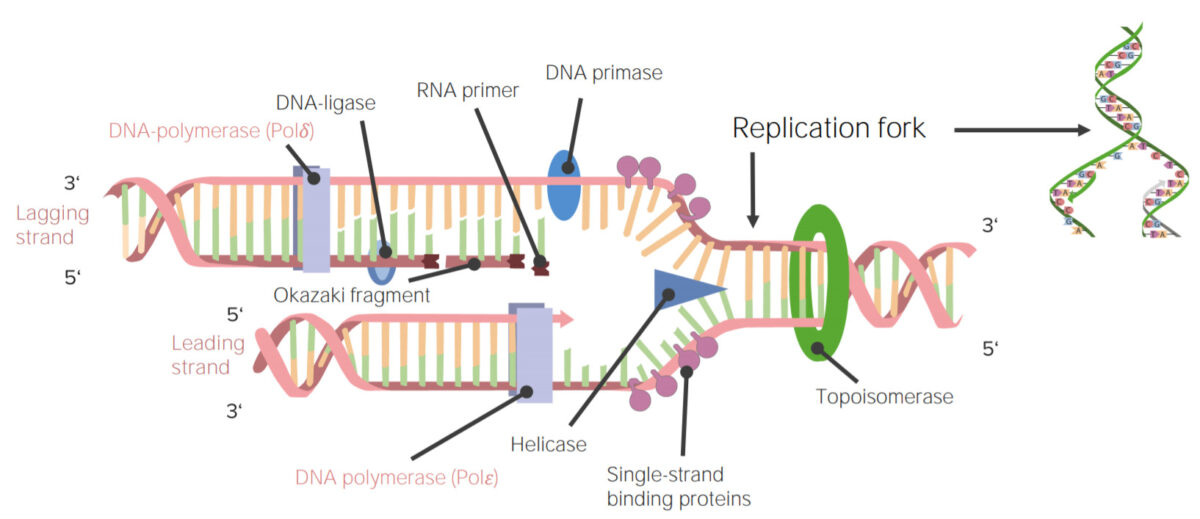
DNA replication or DNA synthesis is the process of copying a double-stranded DNA molecule. This process is paramount to all life as we know it.
Image: “DNA Replication” by Mariana Ruiz. License: Public Domain| Polymerase | Function | Exonuclease activity |
|---|---|---|
| α | Synthesizes the RNA RNA A polynucleotide consisting essentially of chains with a repeating backbone of phosphate and ribose units to which nitrogenous bases are attached. RNA is unique among biological macromolecules in that it can encode genetic information, serve as an abundant structural component of cells, and also possesses catalytic activity. RNA Types and Structure primer and initiates DNA DNA A deoxyribonucleotide polymer that is the primary genetic material of all cells. Eukaryotic and prokaryotic organisms normally contain DNA in a double-stranded state, yet several important biological processes transiently involve single-stranded regions. DNA, which consists of a polysugar-phosphate backbone possessing projections of purines (adenine and guanine) and pyrimidines (thymine and cytosine), forms a double helix that is held together by hydrogen bonds between these purines and pyrimidines (adenine to thymine and guanine to cytosine). DNA Types and Structure synthesis Synthesis Polymerase Chain Reaction (PCR) along the lagging strand | 3′ → 5′ |
| β | Repairs DNA DNA A deoxyribonucleotide polymer that is the primary genetic material of all cells. Eukaryotic and prokaryotic organisms normally contain DNA in a double-stranded state, yet several important biological processes transiently involve single-stranded regions. DNA, which consists of a polysugar-phosphate backbone possessing projections of purines (adenine and guanine) and pyrimidines (thymine and cytosine), forms a double helix that is held together by hydrogen bonds between these purines and pyrimidines (adenine to thymine and guanine to cytosine). DNA Types and Structure | None |
| γ | Replicates mitochondrial DNA Mitochondrial DNA Double-stranded DNA of mitochondria. In eukaryotes, the mitochondrial genome is circular and codes for ribosomal rnas, transfer rnas, and about 10 proteins. DNA Types and Structure | 3′ → 5′ |
| δ | Synthesizes the lagging strand, filling DNA DNA A deoxyribonucleotide polymer that is the primary genetic material of all cells. Eukaryotic and prokaryotic organisms normally contain DNA in a double-stranded state, yet several important biological processes transiently involve single-stranded regions. DNA, which consists of a polysugar-phosphate backbone possessing projections of purines (adenine and guanine) and pyrimidines (thymine and cytosine), forms a double helix that is held together by hydrogen bonds between these purines and pyrimidines (adenine to thymine and guanine to cytosine). DNA Types and Structure gaps after removal of primer | 3′ → 5′ |
| ε | Synthesizes the leading strand | 3′ → 5′ and 5′ → 3′ |