Playlist
Show Playlist
Hide Playlist
Humoral Immunity: Structure of Antibodies and Immunoglobulin Domains
-
08 Slides Humoral Immunity.pdf
-
Reference List Immune System.pdf
-
Download Lecture Overview
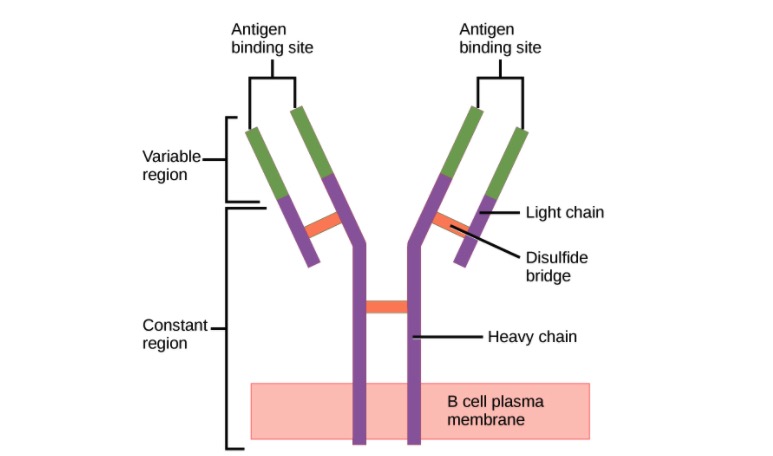
00:01 Let us look at the structure of an antibody molecule. 00:04 They consist of two identical heavy chains, each of which is about 50 kilo Daltons in size. 00:13 And two identical light chains, each of which is around about 25 kilo Daltons in size. 00:21 So adding those together, we can see that the basic antibody molecule is around about 150 kilo Daltons. 00:30 They vary a little bit from class to class, but that basic structure is around about 150 kilo Daltons. 00:37 The two heavy chains are linked to each other by a disulfide bond. 00:42 And each light chain is disulfide bonded to one of the heavy chains. 00:47 This creates a very stable structure. 00:50 Remember disulfide bonds are covalent bonds that are essentially irreversible. 00:59 The antibody can be divided into two different parts - the Variable region and the Constant region. 01:06 So here we can see the Variable region highlighted. 01:11 And this is the part that varies from one antibody to another as the name suggests, and of course that’s related to the specificity of the antibody. 01:19 In other words, to antigen binding. 01:23 In contrast, the Constant regions of the antibody, particularly of the heavy chain are involved in the effector function of the antibody. 01:32 That means what the antibody is actually going to go on and do in order to get rid of the particular infection. 01:39 We can also divide the antibody up into what are called the Fab fragments - the Fragment antigen binding. 01:49 Here, one of the Fragment antigen binding areas is indicated. 01:55 And there are a total of two Fabs. 02:00 And a Fragment crystallizable or Fc fragment which comprises the majority of the Constant region of the heavy chains. So here you can see the two Fabs and the Fc that comprise an antibody molecule. 02:22 The light chain Constant region can either be a kappa light chain Constant region or a Lambda light chain. 02:30 And because the two heavy chains in a single antibody molecule are identical to each other, and the two light chains are identical to each other, if one of the light chains is a lambda light chain, the other will be a lambda light chain. 02:46 If one’s a kappa light chain, the other will be a kappa light chain. 02:49 You never get a mixture of two different types of light chain in a single antibody molecule. 02:59 In terms of the different classes or isotypes of antibodies, this is determined by the heavy chain Constant region. So the mu heavy chain specifies that the antibody is an IgM class of antibody. It’s what’s defining it as an IgM. 03:15 We call an antibody with a C mu Constant region, an IgM antibody. 03:21 If it has a delta Constant region in the heavy chain, that defines it as being an IgD class of antibody. 03:29 Gamma would be IgG, and in fact there are four subclasses of IgG : IgG1, IgG2, IgG3 and IgG4. 03:40 Alpha heavy chain Constant region classifies the antibody as being IgA. 03:46 And there are two subclasses of IgA: IgA1 and IgA2. 03:50 And then finally, if the Constant region of the heavy chain is a C epsilon region, then it’s an IgE antibody. 03:59 The immunoglobulin heavy chain and the immunoglobulin light chain are folded up into domain type structures. 04:09 The light chains have two domains: one Variable domain and one Constant domain. 04:16 Whereas the heavy chains also have a single Variable domain, and then have either three or four Constant domains. An antibody molecule with three Constant domains for the heavy chain is shown in this particular image. And IgG, IgA and IgD antibodies each have three Constant domains in their heavy chain, whereas IgM and IgE antibodies have an additional Constant region domain, making a total of four Constant domains in each heavy chain. The way in which these polypeptide chains are folded, the heavy and the light chain of the antibody molecule can be seen on this diagram. So here we have the Variable domain of the light chain and the Constant domain of the light chain folded into this domain type structure which is stabilized by disulfide bonds within the chain. What we refer to as intra-chain disulfide bonds - within the chain. And the heavy chain here, you can see again, folded up into one Variable domain and three Constant domains. Between the CH1 domain and the CH2 domain, there is an area that is called the hinge. This gives flexibility to the antibody molecule. The two arms of the antibody molecule can move in order to facilitate binding to the antigen. And here you can see the complete antibody molecule with the other heavy chain and the other light chain. And at the tip, the antigen binding site, made up from the VH domain and the VL domain of each antigen binding arm. 06:18 Looking in a little bit more detail at exactly how the protein is folded, we can see that it has this very typical structure. 06:27 And in fact, this is called an immunoglobulin domain type structure. 06:31 And it’s found in many, many different molecules in the body, not just immunoglobulins. 06:36 But it was first discovered in immunoglobulins, which is why it’s called an immunoglobulin type domain. 06:42 And the immunoglobulins remember, is the biochemical term if you like for antibodies. 06:48 So here we have a light chain, the Constant region of the light chain and the Variable region of the light chain. 06:55 Both folded up into this immunoglobulin type domain structure. 07:01 Importantly what you can see here, highlighted in blue, is the three hypervariable regions within the Variable region. 07:11 In other words, the Complementarity Determining Regions or CDRs - CDR1, CDR2 and CDR3. 07:21 And they’re called Complementarity Determining Regions because the amino acids within these CDRs are complementary to amino acids in the antigen that is going to be recognized. 07:33 And illustrated here is the CDR distribution within the light chain Variable region and the heavy chain Variable region. So in total, there are six CDRs in each antigen binding site; three of them coming from the Variable region of the heavy chain and three of them from the Variable region of the light chain. You can see the antigen binding to one arm, and the other arm will bind an identical copy of the antigen. Remember, because the two heavy chains are identical to each other, the two light chains are identical to each other, the antigen binding specificity of the two arms - absolutely identical.
About the Lecture
The lecture Humoral Immunity: Structure of Antibodies and Immunoglobulin Domains by Peter Delves, PhD is from the course Humoral Immunity and Cell-mediated Immunity. It contains the following chapters:
- The Structure of Antibodies
- Immunoglobulin Domains
- Immunoglobulin Domains
Included Quiz Questions
Which of the following immunoglobulin (Ig) antibodies has two subclasses?
- IgA
- IgM
- IgG
- IgD
- IgE
Which of the following is NOT a characteristic of the structure of an immunoglobulin G (IgG) antibody?
- Light chains are attached to heavy chains with hydrogen bonds.
- It consists of two antigen-binding fragment regions and one fragment crystallizable (Fc) region.
- Heavy chains are held together by disulfide bonds.
- Approximate size is 150kDa.
- It has a variable region and a constant region.
Which of the following is NOT an immunoglobulin isotype?
- Kappa
- Mu
- Delta
- Gamma
- Epsilon
Which of the following options correctly pairs the antibody chain with its correct number of constant region domains?
- IgM heavy chains - 4 constant region domains
- Kappa light chains - 2 constant region domains
- IgA heavy chains - 2 constant region domains
- IgE heavy chains - 3 constant region domains
- IgD heavy chains - 4 constant region domains
All EXCEPT which of the following are characteristics of complementarity-determining regions (CDRs)?
- They are present in the constant region of both light and heavy chains.
- They are the most variable parts of the immunoglobulin molecule.
- They have complementary amino acid sequences in the binding antigen
- There are 6 CDRs in each antigen binding site.
- There are 3 CDRs in each variable domain.
Customer reviews
5,0 of 5 stars
| 5 Stars |
|
6 |
| 4 Stars |
|
0 |
| 3 Stars |
|
0 |
| 2 Stars |
|
0 |
| 1 Star |
|
0 |
Very complete and simple to understand!! The presentation has the perfect visual representations.
everything is simple, connected, best explanation and simple slides doctor you are the best thank you
Revision and revision : great efforts ! You just keep on connecting and repeating things that makes everything comprehensible !
the lecture was thorough and used little time to explain and it was very understandable.