Playlist
Show Playlist
Hide Playlist
Pattern Recognition Receptors (PRRs)
-
Slides Innate Immune System.pdf
-
Reference List Immune System.pdf
-
Download Lecture Overview
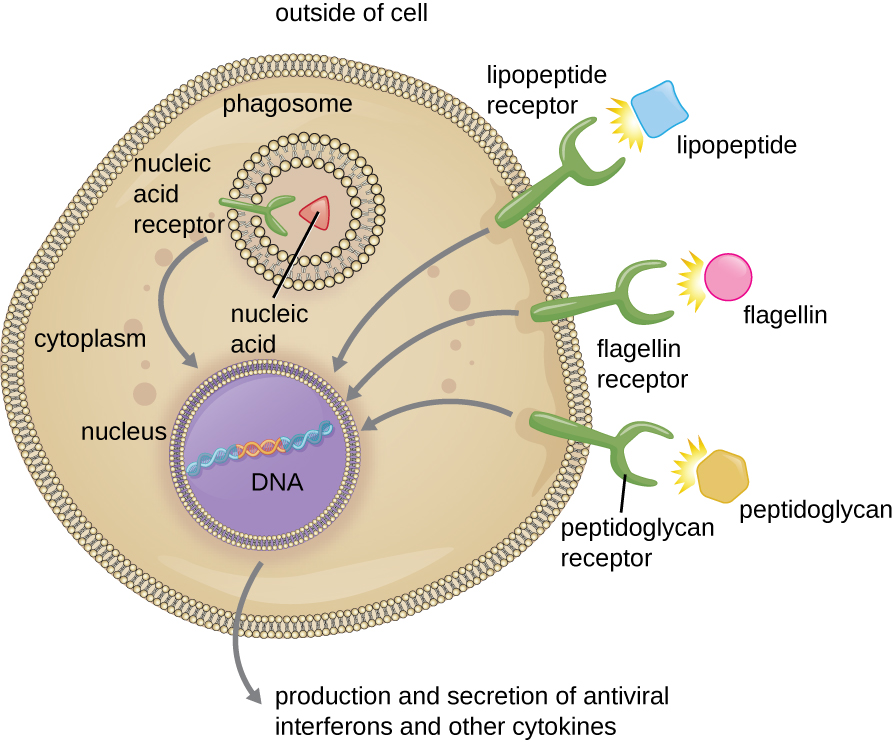
00:01 So how is a threat detected? Well, there are these Pattern Recognition Receptors - PRRs, that can recognize infectious agents. 00:13 And what they recognize on the infectious pathogen are structures that are called Pathogen-Associated Molecular Patterns or PAMPs, P-A-M-P. 00:28 Also, Pattern Recognition Receptors, as well as recognizing PAMPs on foreign infectious agents can recognize structures associated with damage to our own body cells. 00:42 And we call these structures Damage-Associated Molecular Patterns. 00:47 So Pattern Recognition Receptors can recognize both PAMPs and DAMPs. 00:54 And these can be present on the cell surface or sometimes inside cells, and these may be on pathogen cells or they may be on our own body cells. 01:05 What is important to appreciate, is that although this recognition is often described as being broadly specific, what is actually being recognized is recognized in a very, very highly specific way. 01:20 So for example, we’ll mention a few Pathogen-Associated Molecular Patterns in a few moments. 01:26 One of them is called lipopolysaccharide or LPS. 01:31 LPS is found on Gram-negative bacteria. 01:36 And there are lots of different types of Gram-negative bacteria. 01:39 So LPS is shared between several different bacteria but the recognition of LPS is very, very highly specific. 01:48 So recognition is structurally-specific but what is recognized is common to whole groups of organisms or host cells. These Pattern Recognition Receptors can be inside cells, in other words, intracellular. And if they’re intracellular, if they’re inside a cell, they may be present on the endosomes within the cell or they may be present within the cytosol of the cell. Alternatively, they may be present on the surface of cells, cell surface Pattern Recognition Receptors. Or indeed, they may be released or secreted from cells as soluble Pattern Recognition Receptors. You can now look at a number of different Pattern Recognition Receptors and the PAMPs that they recognize. 02:48 Let’s start with endosomal Pattern Recognition Receptors. 02:53 There is a group of 10 or so Pattern Recognition Receptors that are called toll-like receptors. 03:01 Couple of examples for you now; TLR3, toll-like receptor recognizes the PAMP viral double-stranded RNA. 03:14 TLR7 and TLR8 recognize viral single-stranded RNA. 03:22 Whereas TLR9 recognizes a particular nucleotide sequence within the DNA of bacteria, called bacterial unmethylated CpG. 03:36 Let’s now turn to cytosolic Pattern Recognition Receptors. 03:40 NOD-1 (nucleotide-binding oligomerization domain-containing protein-1) and NOD-2. 03:47 These recognize bacterial peptidoglycans. 03:51 These structures are found on Gram-positive bacteria. 03:56 So again, shared between many different bacteria but the recognition of peptidoglycan by NOD-1 and NOD-2 is highly specific for that particular structure. 04:06 RIG-1 (retinoic acid-inducible gene 1) recognizes viral double-stranded RNA. 04:14 And as a third example of a cytosolic Pattern Recognition Receptor, NLRP3 (NOD-like receptor family, pyrin domain-containing 3) which is part of the inflammasome which we’ll discuss in a few seconds. 04:29 This recognizes bacterial lipopolysaccharide (LPS). 04:35 Cell surface Pattern Recognition Receptors; TLR2, again recognizes bacterial structures that are shared between many different bacterial species, various bacterial lipopeptides and lipoproteins. 04:52 TLR4 recognises bacterial lipopolysaccharide (LPS). 04:58 And TLR5 recognises bacterial flagellin. 05:03 And finally, soluble Pattern Recognition Receptors. 05:08 Mannose binding lectin that recognizes the sugar mannose as its name suggests. 05:14 And Ficolin, which recognizes N-acetylglucosamine, another sugar. 05:20 Let’s now turn to Pattern Recognition Receptors which recognize DAMPs - Damage-Associated Molecular Patterns or sometimes called Danger-Associated Molecular Patterns. 05:31 These are Pattern Recognition Receptors that recognize structures produced by our own body cells following damage. 05:41 A couple of examples of Pattern Recognition Receptors present on cell surfaces that recognize DAMPS: RAGE, the receptor for advanced glycation end products as its name suggests recognizes advanced glycation end products that are produced by our own cells in response to damage. 06:03 And RAGE, TLR2 and TLR4 recognize HMGB1 (high mobility group box 1). 06:16 So if you were paying attention you’ll see that RAGE recognizes two different DAMPs - advanced glycation end products and HMGB1. 06:27 Whereas TLR2 and TLR4 are specific not for advanced glycation end products, but for HMGB1. 06:39 And then finally, cytosolic Pattern Recognition Receptors, involved with a structure called the inflammasome; NLRP3 recognizes the Damage or Danger-Associated Molecular Pattern, uric acid.
About the Lecture
The lecture Pattern Recognition Receptors (PRRs) by Peter Delves, PhD is from the course Innate Immune System. It contains the following chapters:
- Pattern Recognition Receptors
- Overview of DAMPs and PAMPs
Included Quiz Questions
Which of the following signals on a bacterial pathogen is mainly recognized by toll-like receptor 4?
- Lipopolysaccharide
- Viral nucleotides
- Uric acid
- Mannose
- Cellulose
Which of the following can be recognized by pattern recognition receptors?
- Damage-associated molecular patterns and pathogen-associated molecular patterns
- Pathogen-associated molecular patterns on host cells
- Damage-associated molecular patterns on pathogens
- Neither damage-associated molecular patterns or pathogen-associated molecular patterns
- Only Pathogen-associated molecular patterns on pathogens
Which of the following is most accurate about the NLRP3 receptor?
- It recognizes the presence of uric acid.
- It is part of the adaptive immune system.
- It recognizes viral single stranded RNA.
- It is only able to recognize damage-associated molecular patterns.
- It is found in the cell's nucleus
Customer reviews
5,0 of 5 stars
| 5 Stars |
|
3 |
| 4 Stars |
|
0 |
| 3 Stars |
|
0 |
| 2 Stars |
|
0 |
| 1 Star |
|
0 |
Excellent lecture, well explained and very concise as always from Dr. P
Gave me a clear informations in simplest way!! Also her accent very God!! i'm from KSA and i found it easily to understand Love it! Thank u.
Great way to explain and communicate the ideas to the students.